
Probing the value of rice grain with premium traits
Advances in rice science have typically veered towards increasing food production to feed a growing population. Rice breeders have been focused on replacing low yielding varieties with higher yielding ones--to produce more grain.
However, these efforts often fail to combine high yields with superior quality grains, leaving farmers with a low-value product and consumers with rice they find unappealing to eat.

Rice quality is primarily assessed based on physical properties such as head rice recovery, chalkiness, grain size and shape, and grain color, and premium quality traits such as aroma have extra value.
Grain quality traits are controlled by major and minor quantitative trait loci (QTLs), implying that the genetic mechanisms underlying quality traits are complex. The most important trends in the research of linking sensory to quality attributes lie in the deeper understanding of the structures, the biochemistry, and the quantities of rice grain components that affect consumer perception.
Eventually, a genetic understanding of sensory attributes can ultimately help breeding programs predict the quality of their newly developed breeding lines at faster rate.
Grain quality relevance to rice breeding
The Grain Quality and Nutrition Center carries out research to unravel importance of various grain quality traits and in addition nutrition value from pre-breeding and breeding material to screen and identify the value addition alleles. In close collaboration with PBGB, we are screening on an average 40,000 lines in a year for grain quality suites.
The availability of a reference rice genome sequence from japonica and resequencing information from indica and several other subspecies of rice allowed rapid identification of the genes responsible for seed dimensions as well as for fragrance in basmati and jasmine rice. Cloned genes responsible for rice grain quality traits as well as fine-mapped QTLs form a strong base for marker assisted selection and genomic selection and offer a tool box in efficient breeding for target markets that are segmented according to grain quality preferences.
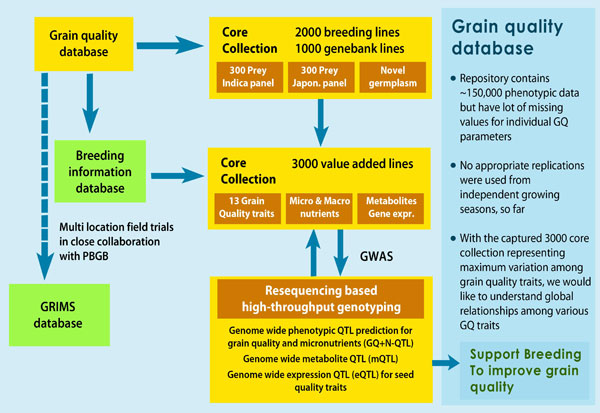
Grain quality genomics
For our research on grain quality and nutrition we utilize re-sequencing data from 3000 lines. The analysis of genomic sequence variations stems from two facts: (1) primarily to use high-density SNP data to clone genes of interest related to grain quality and micronutrients traits, (2) working on structural genomics with a focus on multi-regulatory mechanisms involved in grain quality attributes.
In this context, genome-wide association studies (GWAS) provides a promising tool for the detection of major and minor QTLs and fine mapping of QTLs underlying complex traits of grain quality from untapped germplasm. In this study we will explore the genetic basis of variation for the grain quality and micronutrient traits in a core collection of untapped gene bank germplasm and breeding material.
We have been extensively involved in generating various phenotypic data both on grain quality and micronutrient profiling from this population. By association mapping, we will be able to find both major and minor QTLs associated with various grain quality and micronutrition traits. SNPs reside in those regions, will be extrapolated for marker-assisted selection strategies. In the coming years, subset of lines will be handpicked and transcriptome and metabolome data will be generated from developing seeds. The omics data will be utilized to calculate expression QTLs and metabolite QTLs to identify cis and trans factor in a Systems-Genetics concept.

